In this blog post, I look at whole genome sequence platforms for storage and discuss what might happen to “genomical” amounts of data.
Background
When I uploaded my whole genome sequence in September 2014 (about 10 months ago), few options existed for sharing personal genomic data. The usual suspects (Dropbox, Evernote and Figshare) were prohibitively expensive for large amounts of data. I knew about DNAnexus, but I saw it as a platform for researchers, not consumers. Well, times have changed. Fast.
A Battle of Platforms?
In addition to my original “roll your own” approach, DNAnexus and Google Genomics have emerged as major players for end-to-end genomics workflow. In the table below, you can see that storage costs for AWS S3, DNAnexus and Google Genomics are roughly the same. Everyone provides free uploads (we want your data!), but the cost for transferring data out of the system varies. Google Genomics does not charge for this, but instead charges for API access. For my current AWS storage, I pay about $4 per month to store my genome.
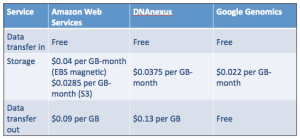
Ultimately, I selected DNAnexus over Google Genomics because their workflow API is well-developed and appealed to my roll-up-your-sleeves sensibility. (If you’re comfortable with command-line work, this platform is for you. BaseSpace, GenoSpace and Galaxy are other platforms to consider.) Google Ventures backed DNAnexus in 2011, so it’s difficult to predict what will happen in the long run. What we do know is that the value of their respective platforms will increase as more people join (and add data) to them. Google Genomics has partnerships with DNAstack, Autism Speaks and even DNAnexus. DNAnexus has partnerships with Baylor College of Medicine, WuXi NextCODE, and the Encode Project. The battle begins. If these two platforms can maintain standards-based interoperability, the competition is good for everyone.
Astronomical becomes Genomical: A Perspective on Storage
In this recent article about big data and genomics, the authors compare the field of genomics with three other Big Data applications: astronomy, YouTube and Twitter. In common with genomics, these domains: 1) generate large amounts of data, and 2) share similar data life cycles. The authors examine four areas–acquisition, storage, distribution, analysis–and conclude that genomics is “on par with or the most demanding” of these disciplines/applications. My previous experience in medical imaging (a field that arguably tackled the prior generation of “big data” issues) leads me to believe that genomics will come to epitomize Big Data to many more people before long.
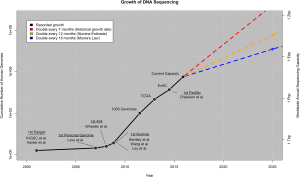
If you look carefully at the projections in the figure above, we may run out of genomes to sequence (really?), which brings us back to storage. Where will we store all of this sequence data, especially as genomic medicine continues its inexorable move to the clinic?
Delete Nothing and Carry on
If the field of medical imaging is an indicator, deleting anything after it has been archived is the exception rather than the rule. The main reason for this is medicolegal — hospitals avoid the liability of not being able to recall an exam later by keeping everything. Although the incidence of requiring access to images after diagnosis is low, the consequence of not having access to the original diagnostic image is high. A former colleague suggested that about 5% of their medical archive customers use lifecycle management features to delete imaging exams. In medical imaging, customers more commonly use lifecycle management features to migrate images to less expensive storage devices over time. So, in genomics, you might migrate your sequence data stored on Amazon from solid state storage (most expensive) to S3 to Glacier (least expensive). But my best guess: we’ll delete nothing and carry on.
Storage is one aspect of genome informatics that is undergoing rapid change. You can learn more at upcoming events like the HL7 2015 Genomics Policy Conference and CSHL’s 2015 Genome Infomatics Conference in October.
Stay tuned!
Update: Why I moved our WGS data from DNAnexus to Amazon S3
Read the updated blog post
